|
10/13/2023 0 Comments 10x chromium droplet basedResearchers then use the barcode tags to identify which bits of RNA or DNA belong to each cell. All the genetic material inside the droplets is then pooled together and sequenced. The cell is then broken apart inside the droplet and the barcode within the bead gets released and attaches itself to the genetic material extracted from the cell. However, advances in technology have led to new methods that can extract and analyse open chromatin or RNA from individual cells.įirst, the cells are separated, via a technique called microfluidics, into tiny droplets of water along with a single bead that carries a unique barcode. Initially, most sequencing tools could only provide an ‘averaged-out’ profile of the genes activated in bulk pieces of tissue which contain multiple types of cell. They do this by extracting and analysing open chromatin (regions of DNA that are accessible to the cell’s machinery), or sequences of RNA (the molecular templates cells use to translate genes into working proteins). These sequencing tools can be used to identify which genes are active within a biological sample. Scientists are now able to determine the order of chemical blocks, or nucleic acids, that make up the genetic code. HyDrop is currently capable of generating single-cell data in high throughput and at a reduced cost compared to commercial methods, and we envision that HyDrop can be further developed to be compatible with novel (multi) omics protocols. Finally, we leveraged HyDrop-RNA’s high capture rate to analyze a small population of fluorescence-activated cell sorted neurons from the Drosophila brain, confirming the protocol’s applicability to low input samples and small cells. Similarly, we applied HyDrop-RNA to flash-frozen mouse cortex and generated 9508 single-cell transcriptomes closely matching reference single-cell gene expression data. 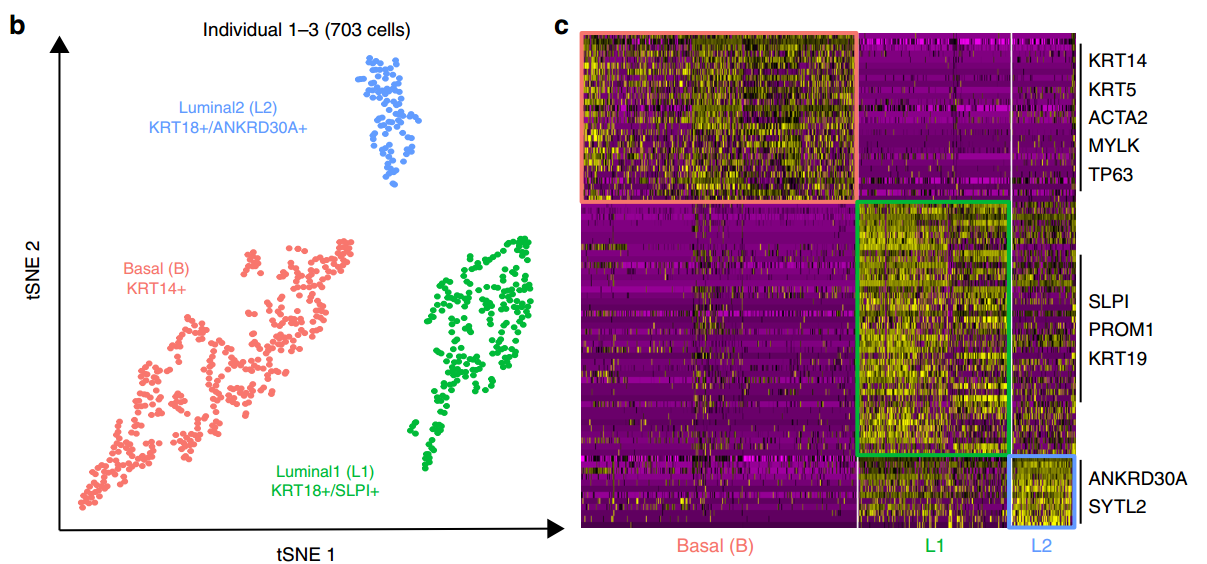
In the third protocol, we adapt both the reaction chemistry and the capture sequence of the barcoded hydrogel bead to capture mRNA, and demonstrate a significant improvement in throughput and sensitivity compared to previous open-source droplet-based scRNA-seq assays (Drop-seq and inDrop). After validating HyDrop-ATAC, we applied it to flash-frozen mouse cortex and generated 7996 high-quality single-cell chromatin accessibility profiles in a single run. In the second protocol, we demonstrate the use of these beads for HyDrop-ATAC, a low-cost noncommercial scATAC-seq protocol in droplets. 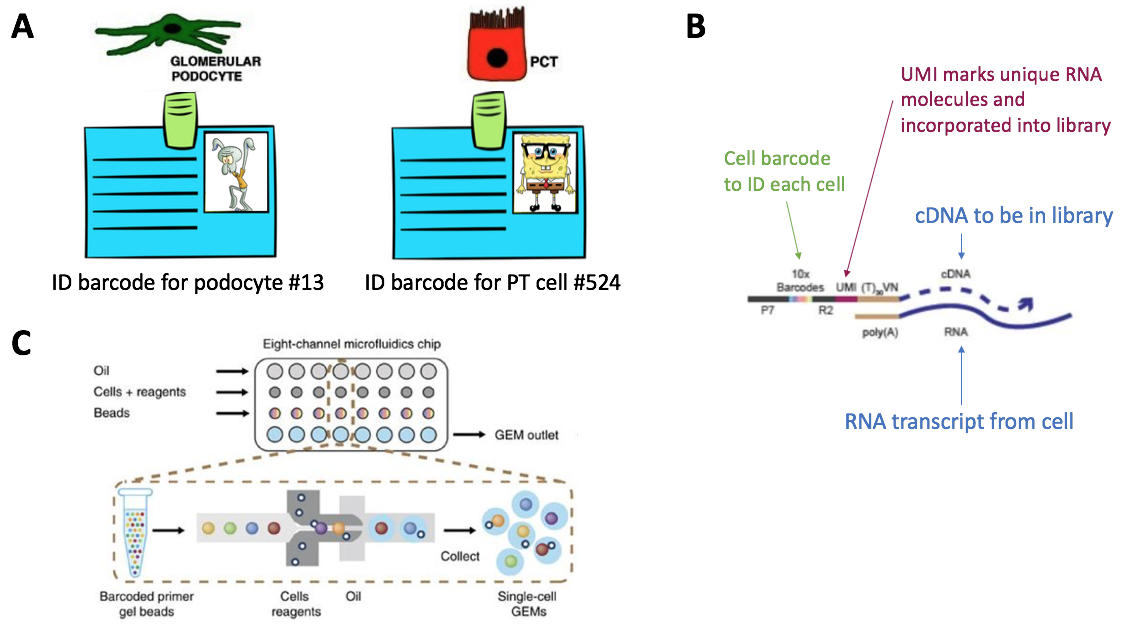 
The first protocol involves creating dissolvable hydrogel beads with custom oligos that can be released in the droplets. We developed HyDrop, a flexible and open-source droplet microfluidic platform encompassing three protocols. To increase the scale of these atlases, lower the cost and pave the way for more specialized multiome assays, custom droplet microfluidics may provide solutions complementary to commercial setups. Single-cell RNA-seq and single-cell assay for transposase-accessible chromatin (ATAC-seq) technologies are used extensively to create cell type atlases for a wide range of organisms, tissues, and disease processes.
0 Comments
Leave a Reply. |
AuthorWrite something about yourself. No need to be fancy, just an overview. ArchivesCategories |
 RSS Feed
RSS Feed
